Difference between revisions of "AutoMeKin"
(→Program execution) |
(→Program execution) |
||
| Line 70: | Line 70: | ||
==Program execution== | ==Program execution== | ||
| − | Unless you donwloaded the singularity image (in that case skip this), to start using any of the scripts of the program, load the amk/20XX module: | + | Unless you donwloaded the singularity image (in that case skip this step), to start using any of the scripts of the program, load the amk/20XX module: |
<code>module load amk/20XX</code> | <code>module load amk/20XX</code> | ||
Revision as of 11:44, 10 February 2020
Contents
Introduction
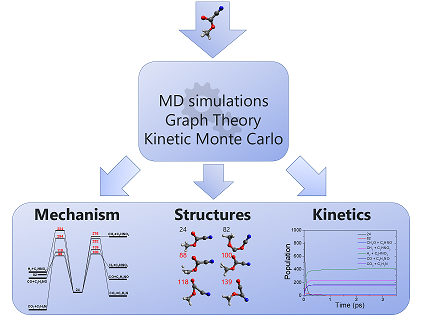
AutoMeKin (formerly tsscds) has been designed to discover reaction mechanisms in an automated fashion. Transition states are located using MD simulations and Graph Theory algorithms. Monte Carlo simulations afford kinetic results. The only input is a starting structure in XYZ format. The method is described in these two publications: 1 2. At present MOPAC2016 and Gaussian 09 (G09) are interfaced with AutoMeKin. The program has been tested on the following Linux distros: CentOS 7, Red Hat Enterprise Linux and Ubuntu 16.04.3 LTS.
To give you a flavor of the capabilities of the program you can try our web interface
Authors
George L. Barnes, David R. Glowacki, Sabine Kopec, Emilio Martinez-Nunez, Daniel Pelaez-Ruiz, Aurelio Rodriguez, Roberto Rodriguez-Fernandez, Robin J. Shannon, James J. P. Stewart, Pablo G. Tahoces and Saulo A. Vazquez
Address
Departamento de Química Física, Facultade de Química, Avda. das Ciencias s/n, 15782 Santiago de Compostela, SPAIN
email me
License
MIT License
Copyright (C) 2020 George L. Barnes, David R. Glowacki, Sabine Kopec, Emilio Martinez-Nunez, Daniel Pelaez-Ruiz, Aurelio Rodriguez, oberto Rodriguez-Fernandez, Robin J. Shannon, James J. P. Stewart, Pablo G. Tahoces and Saulo A. Vazquez
Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
Downloads
2020 version
Download tutorial2020
Options to install/download AutoMeKin2020:
1) This version has been packaged with singularity and the image can be obtained using:
singularity pull library://emartineznunez/default/automekin2020:872
2) Download the tarball from here
3) Clone it from:
90px
2018 version
Download tutorial2018
Options to install/download AutoMeKin2020:
1) Download the tarball from here
2) Cloned it from:
90px
Install the code
To install the code read the Installation instructions
Program execution
Unless you donwloaded the singularity image (in that case skip this step), to start using any of the scripts of the program, load the amk/20XX module:
module load amk/20XX
To run the low-level calculations use:
nohup llcalcs.sh molecule.dat ntasks niter runningtasks >llcalcs.log 2>&1 &
where:
molecule is the name of your molecule
ntasks is the number of tasks
niter is the number of iterations
runningtasks is the number of simultaneous tasks
To run the high-level calculations use:
nohup hlcalcs.sh molecule.dat runningtasks >hlcalcs.log 2>&1 &
For more details, follow the instructions given in the tutorial
References and citations
These four publications must be cited in any work presenting results obtained with our software:
- E. Martínez-Núñez J. Comput. Chem. 2015, 36, 222
- E. Martínez-Núñez Phys. Chem. Chem. Phys. 2015,17, 14912
- A. Rodriguez, R. Rodriguez-Fernandez, S.A. Vazquez, G.L. Barnes, J.J.P. Stewart, E Martínez-Nuñez, J. Comput. Chem. 2018, 39, 1922
- J. J. P. Stewart, MOPAC2016, Stewart Computational Chemistry: Colorado Springs, CO, USA, HTTP://OpenMOPAC.net, 2016.
Publications using AutoMeKin:
- A. Esteban et al. Tetrahedron (2019), doi: https://doi.org/10.1016/j.tet.2019.130764
- R. A. Jara-Toro et al. ChemSystemsChem doi: 10.1002/syst.201900024
- R. Panades-Barrueta et al. Frontiers in Chemistry 2019, 7, 576
- S. Kopec et al. Int. J. Quantum Chem. 2019, 119, e26008
- V. Macaluso et al. J. Phys. Chem. A 2019, 123, 3685-3696
- S. A. Vázquez et al. Molecules 2018, 23, 3156
- A. Rodríguez et al. J. Comput. Chem. 2018, 39, 1922-1930
- D. Ferro-Costas et al. J. Phys. Chem. A 2018, 122, 4790-4800
- Y. Fenard et al. Combust. Flame. 2018, 191, 252-269
- M. J. Wilhelm et al. ApJ. 2017, 849, 15
- J. A. Varela et al. Chem. Sci 2017, 8, 3843-3851
- E. Rossich-Molina et al. Phys. Chem. Chem. Phys. 2016, 18, 22712-22718
- R. Pérez-Soto et al. Phys. Chem. Chem. Phys. 2016, 18, 5019-5026
- S. A. Vázquez and E. Martínez-Núñez, Phys. Chem. Chem. Phys. 2015, 17, 6948-6955
Changelog
Consult the Latest changes
Web interface
AutoMeKin can be used through our web interface.
News
The improvements briefly described below are available in AutoMeKin2020.
AutoMeKin has been interfaced with BXDE to enhance its efficiency ( R. A. Jara-Toro et al. ChemSystemsChem doi: 10.1002/syst.201900024).
The method has also been recently generalized in a collaboration with Dani Pelaez and co-workers to study van der Waals structures ( S. Kopec et al. Int. J. Quantum Chem. 2019, 119, e26008 ) and also to generate sum-of-products PESs for quantum dynamics ( R. Panades-Barrueta et al. Frontiers in Chemistry 2019, 7, 576).
Return to Contents
Return to Main_Page